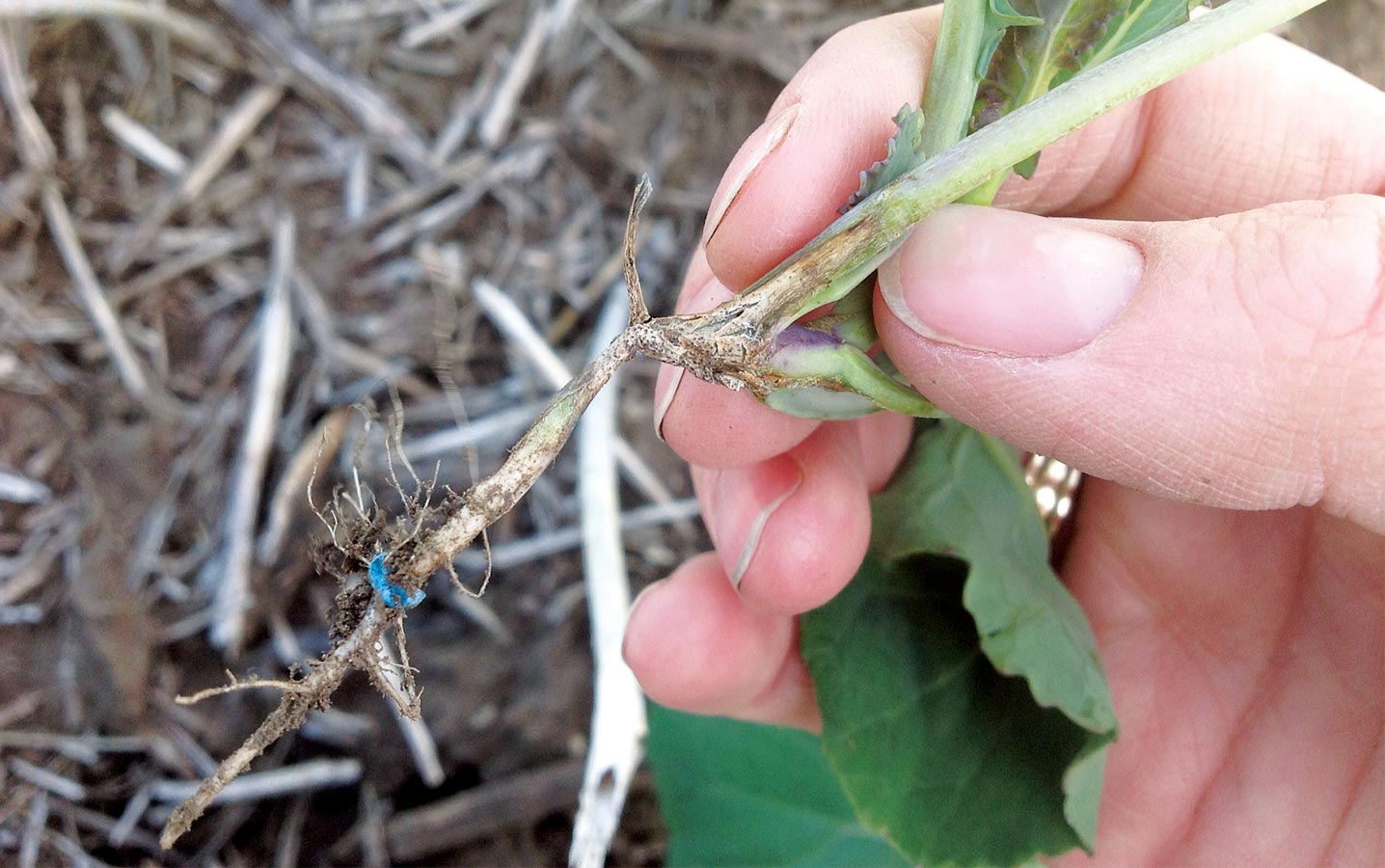
Diversity key to blackleg resistance stewardship
Key practice: Diversity of cultivar resistance, crop rotation and fungicide usage can prevent both infection and breakdown of blackleg resistance.
Project title, Lead researcher: “Blackleg Resistance Stewardship: Improving our management of host resistance,” 2010-14, Dilantha Fernando, University of Manitoba
Grower organization funder: ACPC, MCGA, SaskCanola
Leptosphaeria maculans, the fungal pathogen that causes blackleg in canola, is a limiting factor of canola production in Western Canada. Although resistant cultivars are widely used to control this disease, the breakdown of canola’s genetic resistance, the emergence of new blackleg races, market restrictions, and economic losses are increasing and important concerns to growers.
The objectives of this study, led by Dilantha Fernando, University of Manitoba Department of Plant Science, were to:
- understand pathogen races on-farm and by region;
- identify blackleg-resistant genes in commercial cultivars and strategies to reduce resistance breakdown; and
- initiate a new blackleg management stewardship program that would optimize grower success with moderately-resistant and resistant cultivars through rotations designed to maintain genetic resistance.
Between 2010 and 2012, a total of 930 L. maculans isolates were collected from field stubble across the Prairies and examined for the presence of 10 different avirulence genes. When these genes are absent, there is the possibility of infection on the corresponding blackleg resistance genes in the host. Summary of these results showed that, on average, three of the avirulence genes were absent in more than 90 percent of the pathogen population (AvrLm3, AvrLm9 and AvrLepR2), but two others (AvrLm6 and AvrLm7) were present in more than 85 percent of the population. Blackleg races are composed of combinations of these avirulence genes. In 2010-11, 55 races were detected with each isolate carrying an average of 4.18 out of the
10 genes examined.
Evaluation of 206 Canadian canola lines found the Rlm3 gene to be present in the majority, but all other known Rlm genes were rarely detected. Rlm3 corresponds with the AvrLm3 avirulent gene, which was among the rarest to
be found in all tested pathogen isolates. The low frequency of the AvrLm3 gene coupled with the overuse of Rlm3 single resistance gene to blackleg in planted Canadian canola varieties is the probable cause of current resistance breakdown in many growing regions.
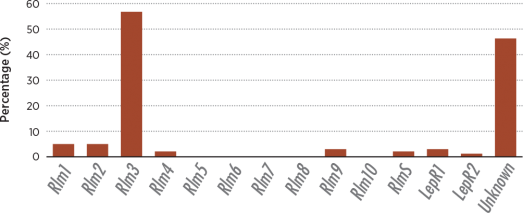
Among the canola varieties examined, 85 percent showed seedling resistance and eight known Rlm genes were found to be present. Rlm3, the most common, was found in 57 percent of the samples. Single-gene resistance was found in 36 percent of these canola samples with the Rlm3 gene again being the clear majority at 92 percent of that subset. All other Rlm traits combined were found in only 14 percent of the samples, and a surprising 43 percent showed no Rlm genes at all.
Among the tested L. maculans isolates, only 8.15 percent carried the AvrLm3 allele, which would prove to be the lowest risk of blackleg infection to canola cultivars carrying this most common Rlm3 gene. More than half of the isolates carried either four or five of the 10 avirulent alleles and a further of the 10 avirulent alleles and a further 11 percent carried three of them. Indications from these findings are that L. maculans diversity is not evenly distributed in Western Canada. Pathogen populations consist of dozens of races and differ greatly from field to field.
Conclusion
Blackleg management recommendations based on this study are:
- Selecting canola varieties with genetic resistance is the best strategy to control blackleg.
- Frequent monitoring of the Avr gene frequency in fungal populations and presence of Rlm genes in canola varieties is critical to disease control.
- Diversification of blackleg resistance in commercial cultivars will allow strategies for disease control such as rotation of diverse Rlm genes.
- The results of this and earlier studies have shown that selection pressure against some Rlm genes may alter the frequency of avirulent genes in L. maculans. This could be exploited for future deployment of Rlm traits.
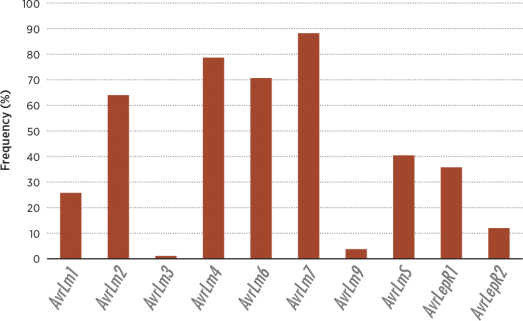
As shown in the figure above, the blackleg fungal population has a high level of diversity in Prairie fields. However, the figure below shows that genetic resistance relies primarily on just one gene, Rlm3. The low frequency of the AvrLm3 allele in the fungal population coupled with the overuse of Rlm3 single resistance to blackleg in planted Canadian canola varieties is the probable cause of current breakdown in many growing regions.