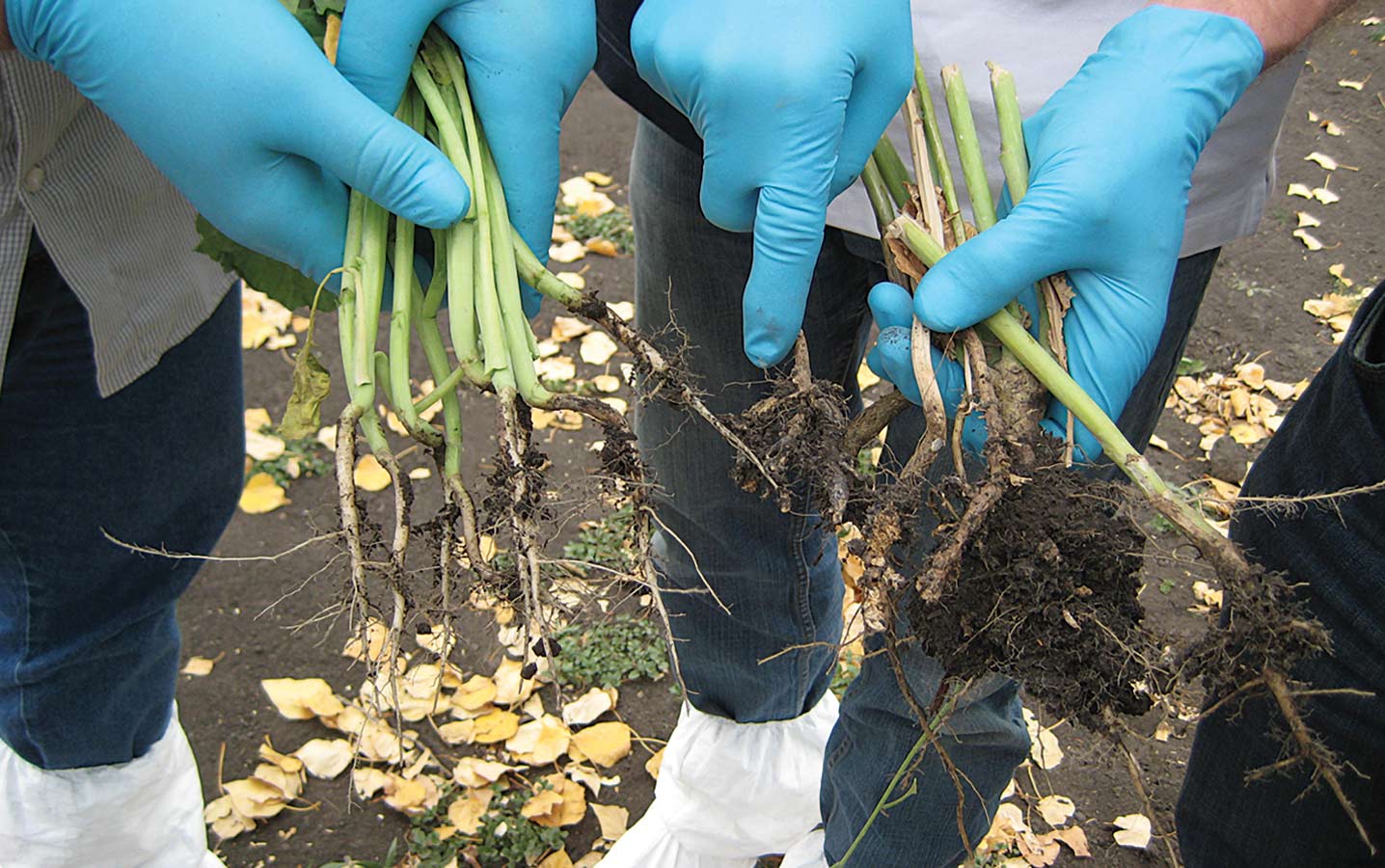
Mapping a route to clubroot resistance
Key practice: P. brassicae can rapidly adapt to the selection pressure provided by currently available clubroot-resistant canola varieties. For durable clubroot resistance, it will be important to stack resistance genes and rotate them in clubroot-infested fields.
Project title, Lead researcher: “Studies on the genetic and molecular basis for clubroot resistance in canola,” 2010-15, Stephen Strelkov, University of Alberta
Grower organization funder: ACPC
The clubroot pathogen Plasmodiophora brassicae has a complex pathotype structure and its soil inoculum levels build up quickly in the presence of susceptible hosts. This means it can quickly adapt to selection pressure. This project provides significant insights to facilitate the improvement and development of sustainable clubroot management strategies.
Project objectives were to:
- identify major resistance genes and develop knowledge of their effective-ness against the predominant pathotypes of P. brassicae in Western Canada;
- gain knowledge of the relationship between different sources of clubroot resistance to enable rotation of resistance genes in infested fields;
- improve understanding of the basis for resistance and influence of different resistance sources on P. brassicae populations in the soil; and
- develop an understanding of a subset of host and pathogen genes that could serve as targets for resistance breeding and fungicide development.
In the clubroot resistance tests, a total of 347 successful double-haploid (DH) canola lines were produced from three different parent populations. The first-generation DH lines were tested with single-spore P. brassicae isolates representing all pathotypes of P. brassicae reported in Canada. Results indicate that a single gene most likely controls P. brassicae resistance in these canola populations.
Pathogen cycling experiments involved five successive cycles of inoculation, resting spore extraction and reinoculation on seven host Brassica genotypes. These genotypes exhibited a range of clubroot resistance from susceptible (S), moderately resistant (MR) to resistant (R). The objectives here were to evaluate changes in pathogen virulence and durability of host resistance at each cycle.
Disease levels in two genotypes — a clubroot-resistant commercial hybrid (CV-R) and a moderately-resistant line — increased significantly after the first inoculation but then remained relatively constant throughout subsequent cycles. In the case of CV-R, this may have reflected the presence of both major and minor clubroot resistance genes, demonstrating the erosion of major-gene resistance at first inoculation followed by partial resistance thereafter.
This quick erosion of clubroot resistance is an accurate prediction of what would happen under field conditions and shows that resistance does not seem to be durable on the presently available CR canola varieties. Indeed, in the 2013 and 2014 growing seasons, several fields in Alberta developed higher clubroot levels than expected, despite being sown to clubroot-resistant (CR) varieties.
Testing on both primary and secondary zoospores indicates that their ability to cause infection may be essentially the same, and that secondary zoospores caused earlier secondary infections than those caused by resting spores. Based on this, the research team hypothesizes that P. brassicae uses primary infection to overcome the plant’s resistance to secondary infection.
P. brassicae zoospore tests conducted on canola and ryegrass plants also provided the first report of host infection by secondary zoospores produced by a non-host. Although rates of secondary plasmodia and infection were highest in canola inoculated with zoospores collected from canola, tiny clubs also developed on 16 percent of the canola plants inoculated with those from the ryegrass. The absence of club formation on the ryegrass in these tests also indicates the build-up of secondary resistance during primary infection in that non-host crop.
Through RNA sequencing of the parent populations and both resistant and susceptible DH lines, a subset of genes has been identified as potentially having key roles in clubroot resistance. Twelve molecular markers were identified that will be useful in breeding programs including gene pyramiding for durable clubroot resistance.A new resistance gene was also identified in one of the B. napus populations providing resistance to P. brassicae either in combination with the previously-identified resistance gene (CRa) or alone, and proving to have a stronger effect than CRa in the population tested.
Findings of this study are being used to formulate recommendations for farmers and have laid a foundation for further research activities.
Conclusions
Recommendations based on this research:
- Manage infected fields through crop rotation out of canola combined with the use of resistant varieties. Even small percentages of susceptible plants within a CR crop can reduce the effectiveness of rotations.
- Practice good sanitation to restrict the movement of possibly contaminated material — resting spores are most likely to spread via contaminated soil and infected canola plant parts.
- Due to P. brassicae’s complex race structure, it has the ability to rapidly adapt to the selection pressure provided by currently available CR canola varieties. Therefore, to provide durable clubroot resistance, it will be important to: (1) stack or pyramid resistance genes in canola varieties; and (2) rotate resistance genes in clubroot-infested fields.
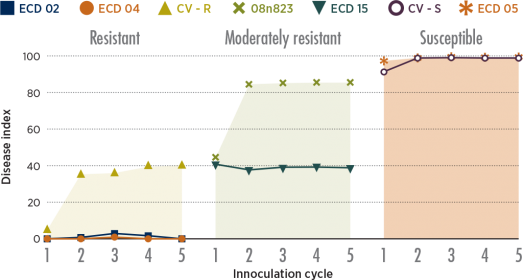
The study put seven Brassica genotypes (denoted by the symbols at the top) through five successive cycles of inoculation, resting spore extraction and reinoculation. Two resistant experimental lines (shown in the graph on the left) held their resistance, with very low levels of disease over the five cycles. Disease levels in the clubroot-resistant commercial hybrid (CV-R) and moderately-resistant line increased significantly after the first inoculation but then remained relatively constant throughout subsequent cycles. This quick erosion of clubroot resistance is an accurate prediction of what would happen under field conditions.