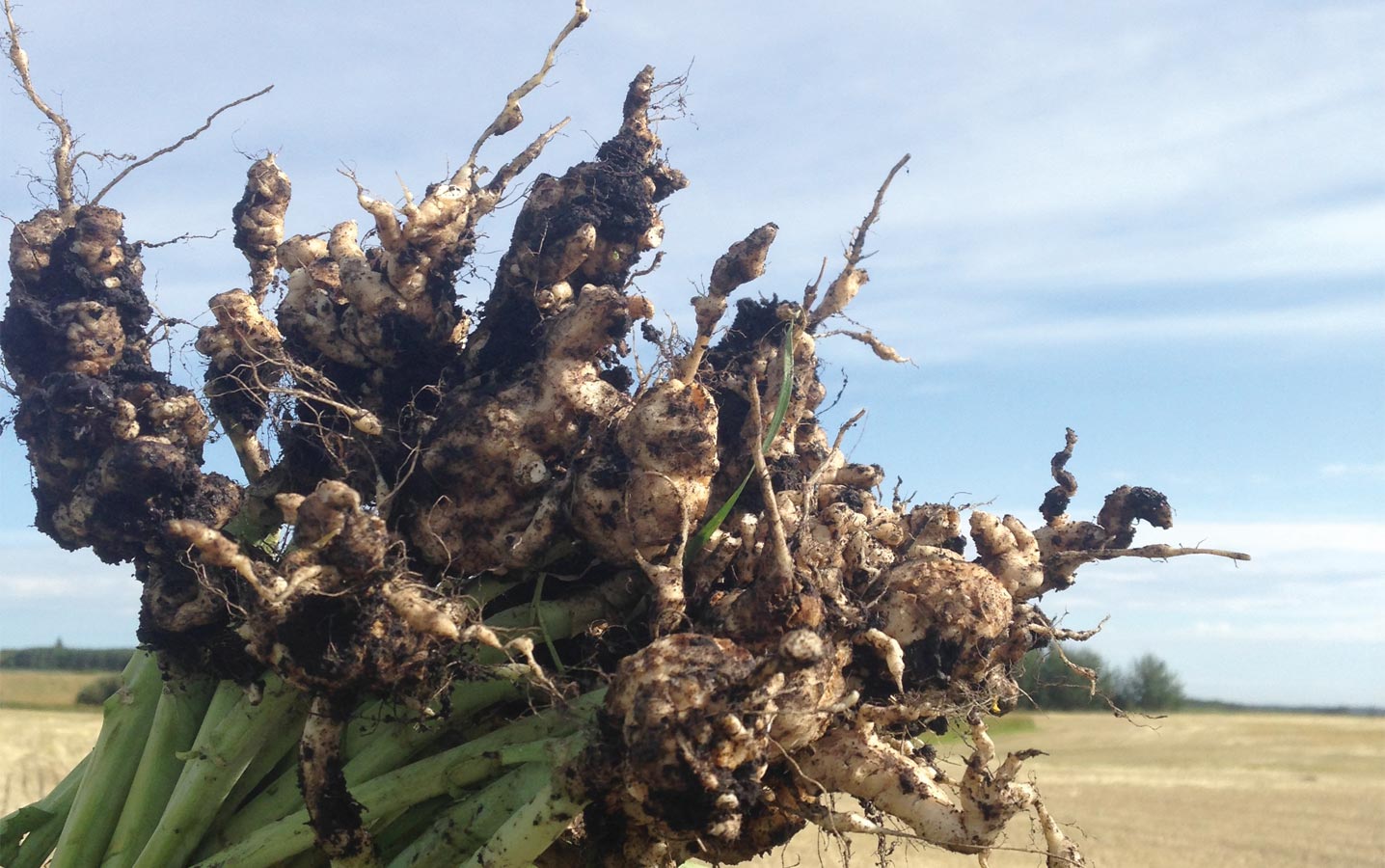
Lots of diversity in clubroot resistance
Key result: Clubroot resistance in Canadian canola varieties relies almost entirely on one source, “Mendel”. This study identified many other genes, found genetic markers for them and then crossed some of them into B. napus lines that could be used for breeding. Canola seed breeders can use the results of this work to broaden and possibly stack the resistance sources they offer.
Project title, Principal investigator: “Identification and mapping of clubroot resistance genes in Brassica and development of SNP markers tightly linked to resistance genes,” Fengqun Yu, Agriculture and Agri-Food Canada, Saskatoon
Funding: GF2, ACIDF, WGRF, Alberta Canola, AAFC
In 24 clubroot-resistant Brassica lines collected from around the world, the type of resistance could be grouped into 16 clusters. The “Mendel” clubroot-
resistance gene, the source resistance for most clubroot cultivars available on the Canadian market, was just one of them.
This is good news given that the effectiveness of Mendel in clubroot-infested parts of Alberta seems to be eroding. Pyramiding of resistance genes or rotating to other sources of resistance could improve clubroot management through genetic resistance.
A previous GF1-funded study by Gary Peng and Kevin Falk at AAFC’s Saskatoon Research Centre identified several B. rapa cultivars/breeding lines and one B. oleracea cultivar highly resistant to five pathotypes
of clubroot-causing P. brassicae previously identified
in western Canada.
In this project, which ran from 2013-16, Yu and her team mapped the genes then identified molecular markers tightly linked to the CR genes in the cultivars that Peng and Falk identified. These DNA markers associated with the target genes, allow breeders to confirm the presence of a desired gene early in the breeding process.
Five resistance genes were mapped in B. rapa: Rcr1 was identified from bok choy cultivar “Flower Nabana”, Rcr2 from Chinese cabbage cultivar “Jazz”, Rcr3 from canola B. rapa breeding line 6992, Rcr4 from canola breeding line T19 and Rcr5 from turnip cultivar “Purple Top White Globe”. As well, Rcr6 was identified in B. nigra line PI and Rcr7 was identified in B. oleracea cabbage cultivar “Tekila”.
Published results
Yu published part of the work – work on Rcr1 – in PLoS ONE journal*. The article describes how Yu and her team found 776,200 single nucleotide polymorphisms (SNPs) and 122,200 insertion/deletion (InDels) in resistant cultivars. SNPs and InDels are patterns that could possibly become “markers”. Through time-consuming lab work using well-established tools, fourteen SNP markers in the target region were confirmed to associate with Rcr1. The team then employed nine of these SNP markers for marker-assisted introgression of Rcr1 into B. napus canola from B. rapa, with 100 per cent accuracy.
Similar work was done for each of the other six clubroot-resistant Brassica lines that Peng and Falk had earlier identified.
The next step was to insert these newly-discovery CR genes into commercial species for breeding purposes. They successfully “re-synthesized” a B. napus canola line to contain Rcr3 and Rcr7 genes, which could greatly broaden the genetic base of clubroot resistance in B. napus. The project also re-synthesized both B. juncea and B. carinata with highly resistant genes. This re-synthesis will enable both canola and mustard breeders to rapidly incorporate clubroot resistance into their cultivars.
*Yu F, Zhang, X, Huang Z, Chu M, Song S, Chang A., Deora A, Chen Q, Zhang Y, McGregor L, Falk KC, Gossen BD, McDonald MR,Peng G. Identification of SNP Markers Associated with Clubroot Resistance Gene Rcr1 through Bulked Segregant RNA Sequencing PLoS ONE 11(4): e0153218. doi:10.1371/journal.pone.0153218.